I developed this project for the second group project period of the MSc in Systems Biology at Maastricht University.It was an end-to-end assignment with data pre-processing, feature engineering, training, hyperparameter tuning, and interpretation of the results.

Background
The human brain processes sound through specialized ventral and dorsal streams within the auditory cortex. While computational models like the Wilson-Cowan (WC) firing-rate model have been developed to simulate these dynamics, their ability to replicate real physiological responses remains to be fully validated.
In this project, we developed AuditWC-fMRI-ML, a classification framework designed to compare the predictive capacity of the WC model against functional magnetic resonance imaging (fMRI) data. The goal was to determine if the model accurately simulates the biological processing of spectrotemporal modulations in sound. To achive this, we trained machine learnning (ML) classifiers on both datasets to classify 288 sounds across six categories (speech, voice, animal, music, nature, and tools) and compared their performance across different brain regions.
Our methodology encompasses four main stages:
- Preprocessing: Concatenation of voxels across subjects and feature extraction for the modeling data.
- Model Selection: Screening algorithms for multiclass classification.
- Hyperparameter Tuning: Fine-tuning the best models using GridSearchCV.
- Evaluation: Comparing regional performance using various classification metrics.
An overview of the workflow of this project is shown in the graphical abstract below.

Figure 1.- Graphical abstract of the AuditWC-fMRI-ML workflow.
The main objectives were to:
- Classify 288 diverse sounds using neural signatures from both fMRI and simulated firing rates.
- Compare the performance of model-simulated data against experimental fMRI responses across specific brain regions (A1, R, Slow, and Fast).
- Identify potential refinements for the firing-rate model based on classification discrepancies.
Dataset
This study uses two distinct datasets containing responses to 288 sounds across six categories: speech, voice, animal, music, nature, and tools.
- Experimental (fMRI) Dataset: Neural responses measured in 47,505 voxels from five subjects, distributed across core (A1, R) and belt (Slow, Fast) areas of the auditory cortex.
- Modeling (WC) Dataset: Simulated firing rates in 98 channels for the four regions. This was supplemented with 2D Fourier Transform (2DFT) features (80 temporal and 48 spectral) to better capture sound modulations.
| Region | Voxel Count (fMRI) | Channels (WC Model) |
|---|---|---|
| A1 | 9,511 | 98 |
| R | 8,884 | 98 |
| Slow | 15,539 | 98 |
| Fast | 13,571 | 98 |
Machine Learning Models
We tested four multiclass classifiers using scikit-learn: Random Forest (RF), XGBoost, Support Vector Machine (SVM), and Multilayer Perceptron (MLP). Models were implemented using the One-vs-All strategy and evaluated through 3-fold cross-validation.
The Random Forest algorithm was the top performer for both the modeling and fMRI datasets and was selected for in-depth comparative analysis.
Results of the ML models
The classifiers achieved performance significantly above random chance, demonstrating that both biological and simulated data capture distinguishable sound features. However, the fMRI-trained models consistently outperformed the basic WC model.
Algorithm Comparison (Cross-Validation Accuracy)
| Dataset | RF | SVM | XGBoost | MLP |
|---|---|---|---|---|
| Modeling | 0.55 | 0.52 | 0.51 | 0.49 |
| fMRI | 0.67 | 0.68 | 0.64 | 0.58 |
Feature Integration Impact (Random Forest Test Data)
Integrating 2DFT features into the modeling dataset significantly closed the performance gap with biological fMRI data.
| Metric | fMRI | Modeling | Modeling + 2DFT |
|---|---|---|---|
| Accuracy | 0.74 | 0.62 | 0.69 |
| F1-score | 0.73 | 0.63 | 0.69 |
| MCC | 0.69 | 0.55 | 0.63 |
The ROC curves below show that RF models achieved high classification performance on the fMRI dataset (AUROC>0.85), with performance declining on the modeling dataset but substantially improving when 2DFT features were added, particularly for speech and voice categories (AUROC>0.95).
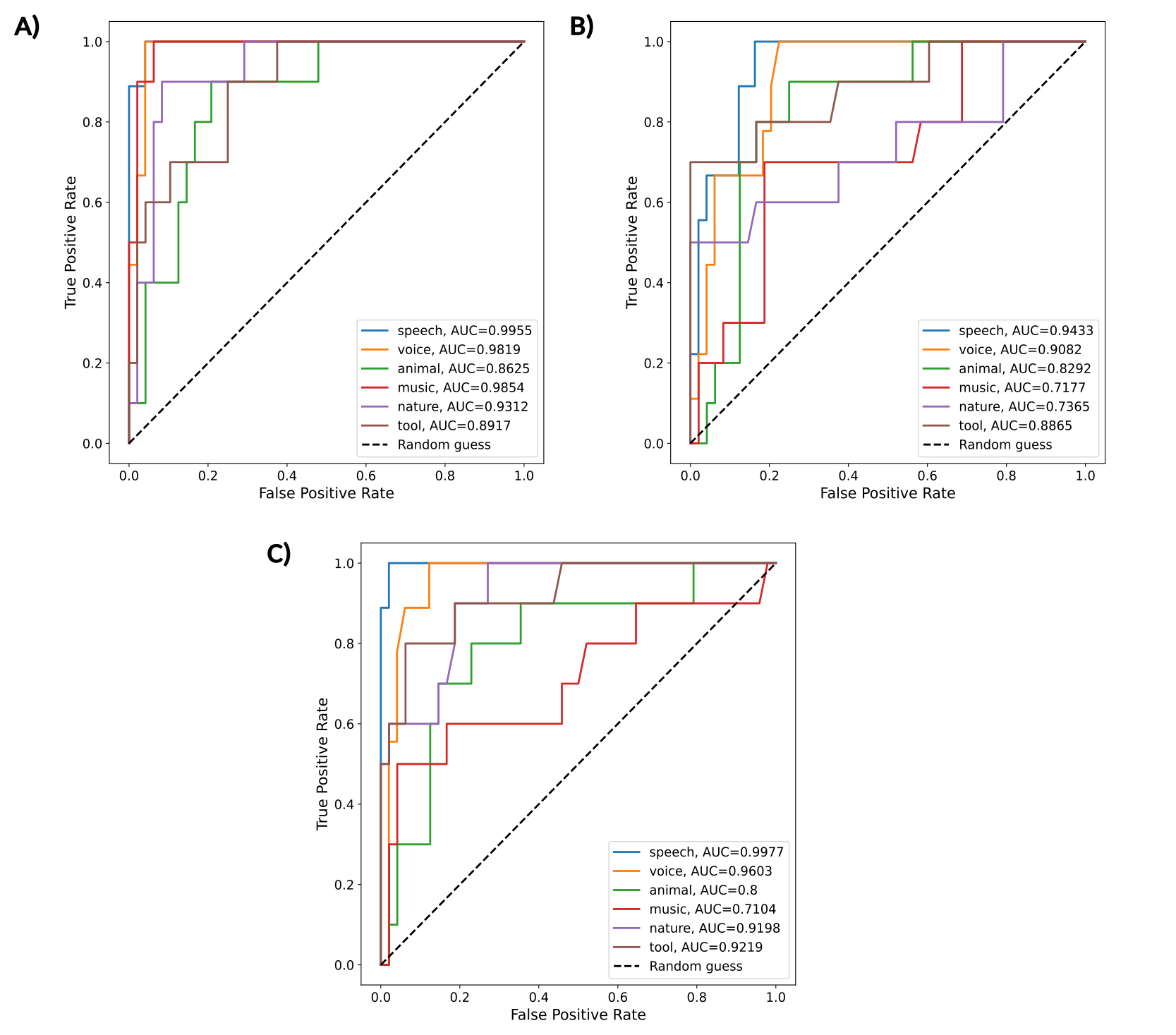
Figure 2.- ROC curves for all the categories of the multi-class Random Forest algorithms for A) fMRI, B) modeling, and C) modeling + 2DFT datasets.
Biological Interpretation
- Regional Specialization: Biological data (fMRI) showed increased classification accuracy in the Slow and Fast regions compared to the primary regions (A1, R), supporting the hypothesis that belt areas are specialized for complex sound characterization.
- Model Limitations: The WC model exhibited uniform performance across all regions, failing to replicate the regional specialization observed in the human brain. This suggests the need for region-specific parameter tuning.
- Information Recovery: The success of the 2DFT features indicates that current firing-rate models lose critical temporal and spectral modulation data that the biological brain preserves.
As shown in the figure below, the fMRI-trained models demonstrate clear regional hierarchy with highest accuracy in the S region, while the basic WC model shows uniform performance across regions—a gap that narrows considerably when 2DFT features are added.
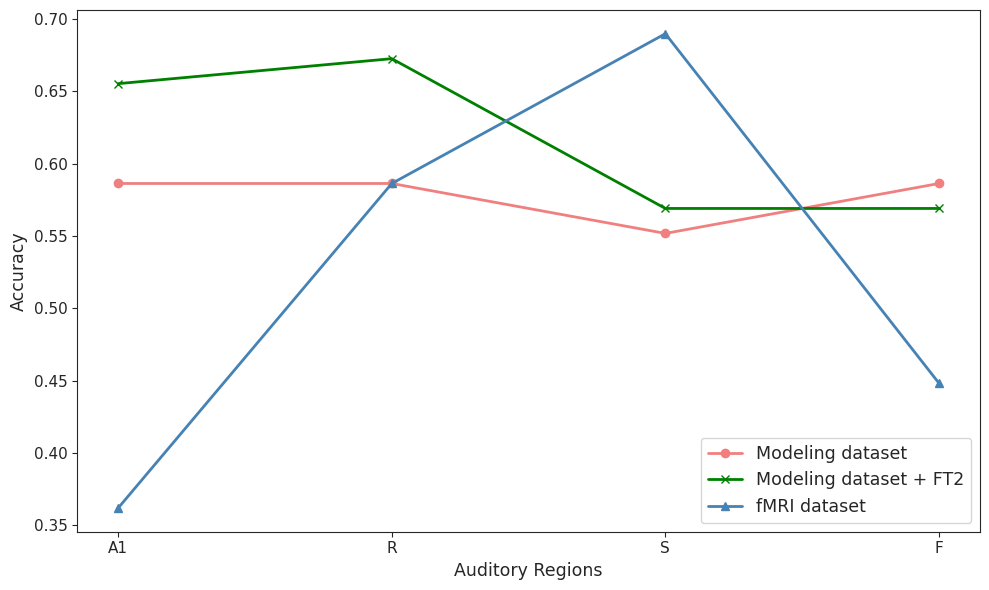
Figure 3.- Accuracy of the classifiers by auditory regions for the fMRI, modeling, and modeling + 2DFT features datasets. Every line represents one dataset and each point corresponds to one auditory region.
Additional information
The complete information regarding the exploratory data analysis and selection of the best model, training and validation python scripts, hypterparameter tuning, and further details are available on the GitHub repository of this project and in the manuscript.