Source code
I developed this project for the first group project period of the MSc in Systems Biology at Maastricht University.It was an end-to-end assignment with data pre-processing, feature engineering, training, hyperparameter tuning, and deployment of the best model as a web application with Streamlit.

Background
Cardiomyopathies are morphological and functional abnormalities in the myocardium that affect millions of people worldwide. Despite a clear genetic component, characterizing the molecular signatures that distinguish different cardiomyopathy etiologies (dilated (DCM), hypertrophic (HCM), or peripartum (PPCM)) remains a significant challenge.
In this project, we developed CardiomyoML, a machine learning framework to classify heart tissue samples based on RNA-seq gene expression data. An overview of the workflow of this project is shown in the graphical abstract below.
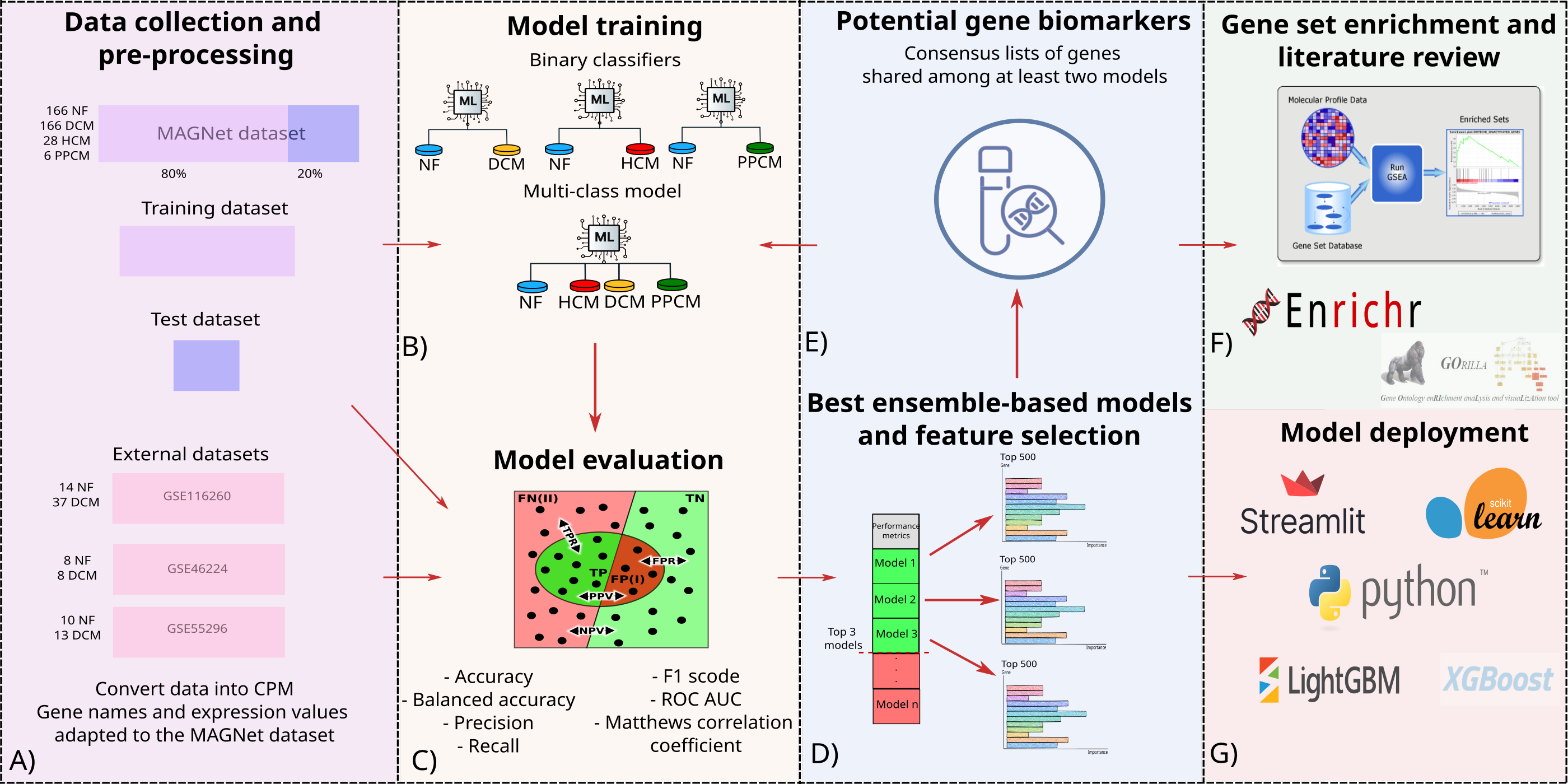
Figure 1.- Graphical abstract of the CardiomyoML workflow.
The main objectives were to:
- Classify samples into Non-failing (NF), DCM, HCM, or PPCM cardiomyopathies.
- Identify a robust set of genetic biomarkers using feature selection and consensus ranking.
- Validate the biological relevance of these genes through enrichment analysis and literature review.
Dataset
The primary dataset for training and testing the ML models was the Myocardial Applied Genomics Network (MAGNet) (GSE141910). For validation, we used three external datasets (GSE46224, GSE116250, and E-GEOD-55296).
| Dataset partition | Samples | NF | DCM | HCM | PPCM |
|---|---|---|---|---|---|
| MAGNet (Training/Test) | 366 | 166 | 166 | 28 | 6 |
| External Validation 1 (GSE46224) | 16 | 8 | 8 | - | - |
| External Validation 2 (GSE116250) | 51 | 14 | 37 | - | - |
| External Validation 3 (GSE55296) | 10 | 13 | - | - | - |
Feature Selection
The feature matrices were generated using log-transformed Counts Per Million (log-CPM) values. To handle the high dimensionality of RNA-seq data, we implemented a multi-stage feature selection process.
- Ensemble Ranking: Extracted the top 500 features from the best-performing ML models.
- Consensus List: Identified a list of 94 genes consistently ranked high across all models.
- Biological Validation: Analyzed consensus genes (e.g., MYH6, NPPA) for Gene Ontology (GO) enrichment.
Machine Learning Models
First, we tested more than 30 ML classifiers for the binary and multiclass classification tasks using the LazyPredict Python library. We chose the top 3 models according to some performance metrics such as accuracy, ROC AUC, precision, recall, F1 score, and Matthews Correlation Coefficient (MCC). Then, we fine-tuned the hyperparameters of the best models using sklearn’s class GridSearchCV. Finally, considering the results of hyperparameter tuning and performance metrics, we obtained the best ML model to predict the cardiomyopathy etiology of the samples.
Results of the best ML models
The models achieved high performance on internal test data, particularly in the NF/DCM task. However, performance significantly decreased when evaluated on external datasets.
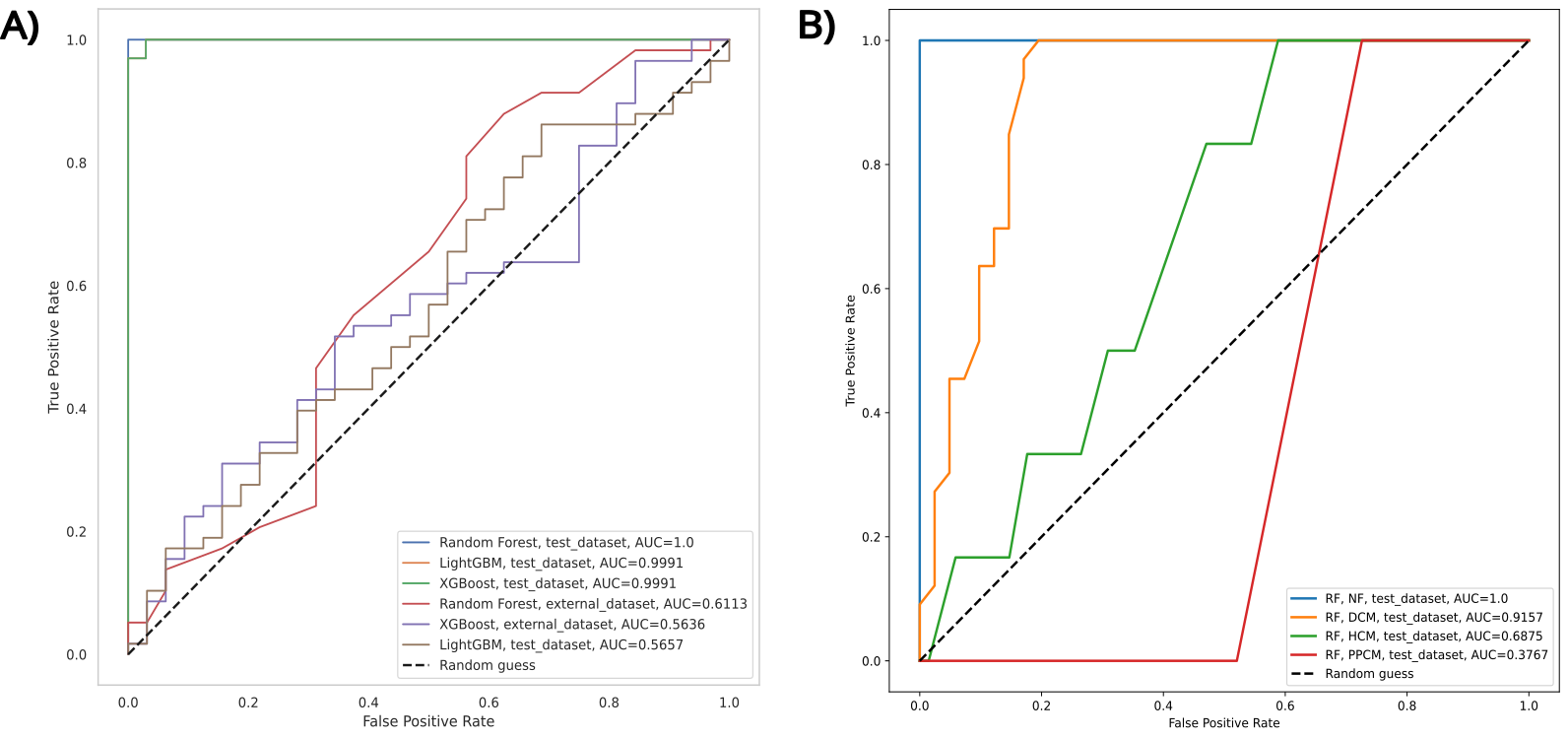
Figure 2. ROC curves of the top 3 ML models in A) the binary classification task of NF/DCM using test and external data and B) individual performances for each etiology in the multiclass classification task on test data.
1. Binary Classification: NF/DCM (Test Data)
| Metric | RF | LGBM | XGBoost |
|---|---|---|---|
| Accuracy | 0.99 | 0.99 | 0.97 |
| Balanced Accuracy | 0.99 | 0.99 | 0.97 |
| Precision | 0.97 | 0.97 | 0.94 |
| Recall | 1.00 | 1.00 | 1.00 |
| F1score | 0.99 | 0.99 | 0.97 |
| MCC | 0.97 | 0.97 | 0.94 |
2. External Validation: NF/DCM (External Data)
| Metric | RF | LGBM | XGBoost |
|---|---|---|---|
| Accuracy | 0.66 | 0.52 | 0.62 |
| Balanced accuracy | 0.58 | 0.48 | 0.55 |
| Precision | 0.69 | 0.63 | 0.68 |
| Recall | 0.84 | 0.62 | 0.79 |
| F1score | 0.76 | 0.63 | 0.73 |
| MCC | 0.18 | -0.04 | 0.12 |
3. Multiclass Classification (Test Data)
| Metric | RF | LGBM | XGBoost |
|---|---|---|---|
| Accuracy | 0.91 | 0.91 | 0.89 |
| Balanced accuracy | 0.50 | 0.53 | 0.49 |
| Precision | 0.83 | 0.91 | 0.82 |
| Recall | 0.91 | 0.91 | 0.89 |
| F1score | 0.86 | 0.88 | 0.85 |
| MCC | 0.84 | 0.84 | 0.82 |
Biological Interpretation
Using the 500 most important genes from the top 3 ensemble models (RF, LGBM, and XGBoost) in the NF/DCM task, we identified 94 consensus genes shared by at least two models, as shown in the Venn diagram in Figure 3A.
- Literature Comparison: When compared against established literature datasets, only one gene, MYH6 (myosin heavy chain 6), was shared. MYH6 is vital for cardiac muscle contraction, and its mutations are linked to cardiomyopathies and sudden cardiac death.
- Enrichment Analysis: GSE analysis using the Enrichr web server identified enriched ontologies related to myocardial infarction and HCM in the Phenotype-Genotype Integrator and OMIM Disease databases (Figure 3B).
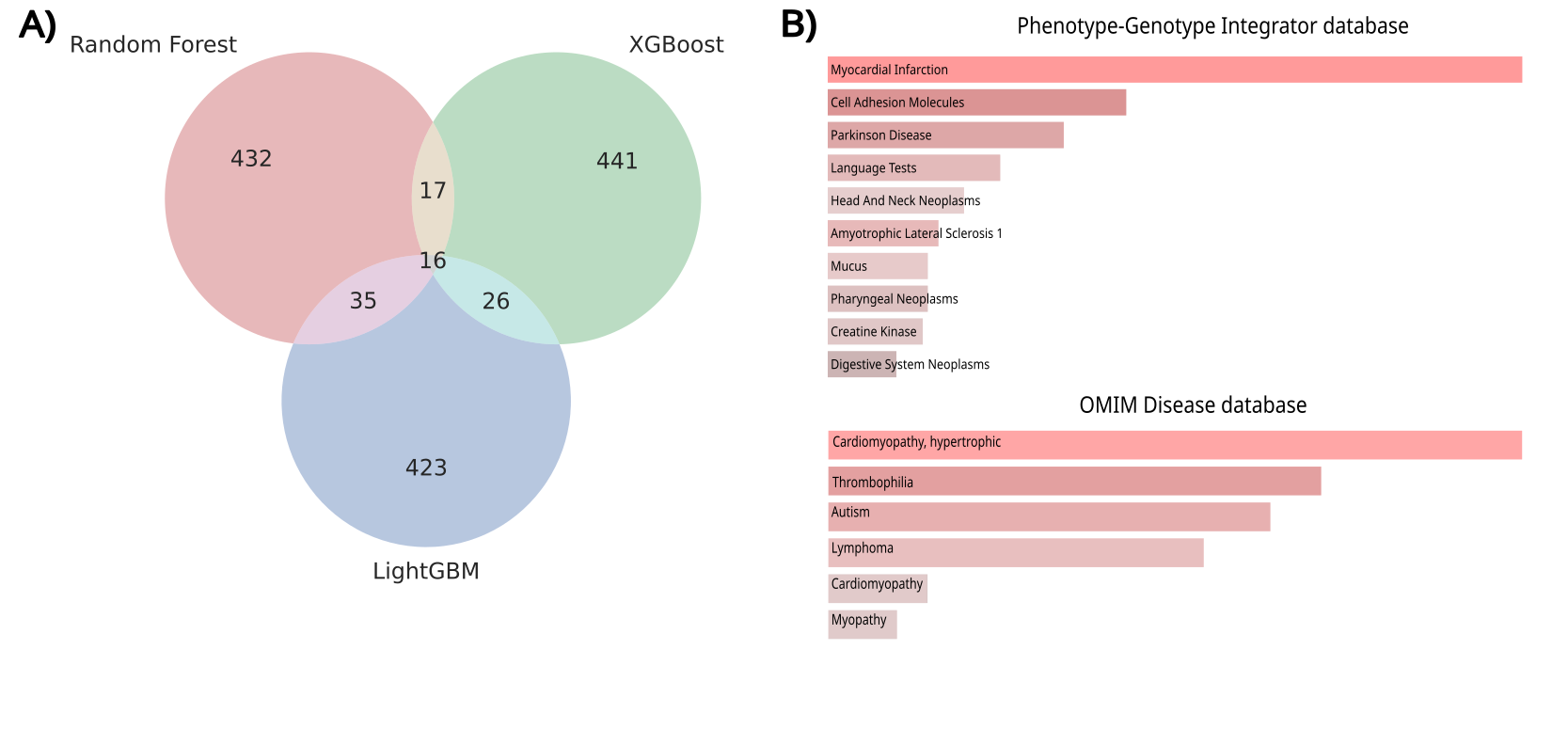
Figure 3. A) Venn diagram of the top 500 most important genes for DCM. B) Gene set enrichment results associated with heart diseases, obtained via Enrichr.
Web Application
Using the best-performing model of the NF vs. DCM task, we developed CardiomyoPred, a Streamlit web application that allows users to predict the cardiomyopathy etiology of heart tissue samples based on gene expression data for the 94 consensus genes. The source code for this web application is available in this GitHub repository.

Additional information
The complete information regarding the exploratory data analysis and selection of the best model, training and validation python scripts, hypterparameter tuning, and further details are available on the GitHub repository of this project and in the manuscript. The supplementary information of this project is available in Zenodo.
