Source code
I developed this project for my capstone project of the Scientific Programming course from the MSc in Systems Biology at Maastricht University. It was an end-to-end assignment with data pre-processing, feature engineering, training, hyperparameter tuning, and interpretation of the results.

Background
Dengue is a viral infection transmitted to humans through the bite of Aedes mosquitoes. This disease is a neglected tropical disease that mainly affects poor populations with no access to safe water, sanitation, and high-quality healthcare. Currently, there is no specific treatment for dengue and the focus is on treating pain symptoms. Therefore, there is an urgent need to find new drugs to treat this disease.
The goal of this project is to predict new repurposed drugs for dengue using a biomedical knowledge graph and graph neural networks. A knowledge graph (KG) is a heterogeneous network with different types of nodes and edges that incorporate semmantic information. A KG is composed of a set of triplets (subject, predicate, object) that represent relationships between entities. For example, a drug-disease triplet represents the relationship between a drug (subject) and a disease (object) through a predicate (e.g., treats, causes, etc.). The advantage of using a KG is that it allows the integration of different types of data from different sources.
Graph neural networks (GNNs) are a class of neural networks that can learn from graph data. GNNs have been used to solve different tasks in KGs, such as node classification, link prediction, and entity alignment. The drug repurposing problem can be formulated as a link prediction task in a KG. The goal is to predict new drug-disease associations for Dengue.
The following figure shows the general workflow of this project:

Figure 1.- Graphical abstract of the DengueDrugRep workflow.
Dataset - Knowledge graph
The DRKG is a large-scale biomedical KG that integrates information from six existing databases: DrugBank, Hetionet, Global network of biomedical relationships (GNBR), String, IntAct, and DGIdb. This KG contains 97.238 nodes belonging to 13 entity-types (e.g., drugs, diseases, genes, etc.) and 5.874.257 triplets belonging to 107 edge-types. Also, the DRKG contains 24.313 compounds from 17 different databases (the list of databases’ names is available in the Names_datasources_compounds_DRKG.csv file).
The following figure shows the metagraph of the DRKG:
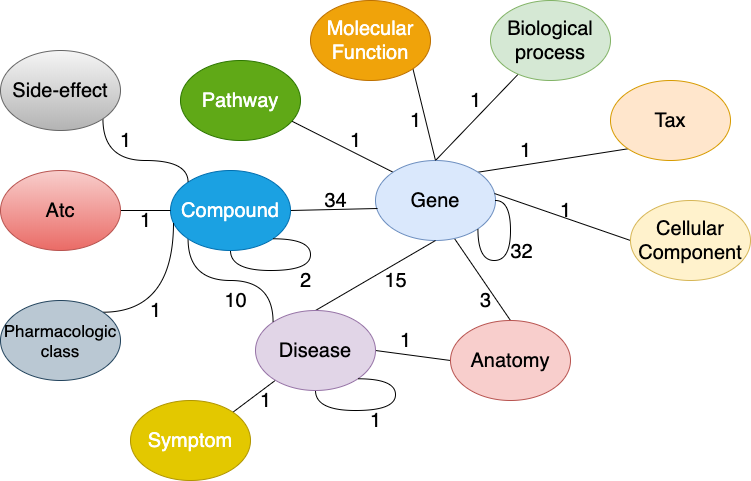
Figure 2.- Metagraph of the DRKG. The number next to an edge indicates the number of relation-types for that entity-pair in the KG. Obtained from [2].
The PyKEEN library implements the DRKG as part of its datasets, so it is possible to load the DRKG directly from the library.
The DRKG was split into training, validation, and test sets. The training set contains 4.699.405 triplets, the validation set contains 587.426 triplets, and the test set contains 587.426 triplets. The training partition was used to train the models, and the validation partition was used to evaluate the models. The test dataset was used to make predictions on unseen data.
Exploratory data analysis
In this project, I focused on the drug-disease relationships in the DRKG. So, the first step was to explore what are the predicates that represent these relationships, obtaining the following list:
| Drug-Disease predicate |
|---|
| DRUGBANK:: treats::Compound:Disease |
| GNBR:: C::Compound:Disease |
| GNBR:: J::Compound:Disease |
| GNBR:: Mp::Compound:Disease |
| GNBR:: Pa::Compound:Disease |
| GNBR:: Pr::Compound:Disease |
| GNBR:: Sa::Compound:Disease |
| GNBR:: T::Compound:Disease |
| Hetionet:: CpD::Compound:Disease |
| Hetionet:: CtD::Compound:Disease |
More details about the predicates, their provenance and meaning are available in the relation_glossary.tsv file and the DRKG GitHub repository.
Next, I explored the number of compounds per database in the DRKG. The following figure shows the results:
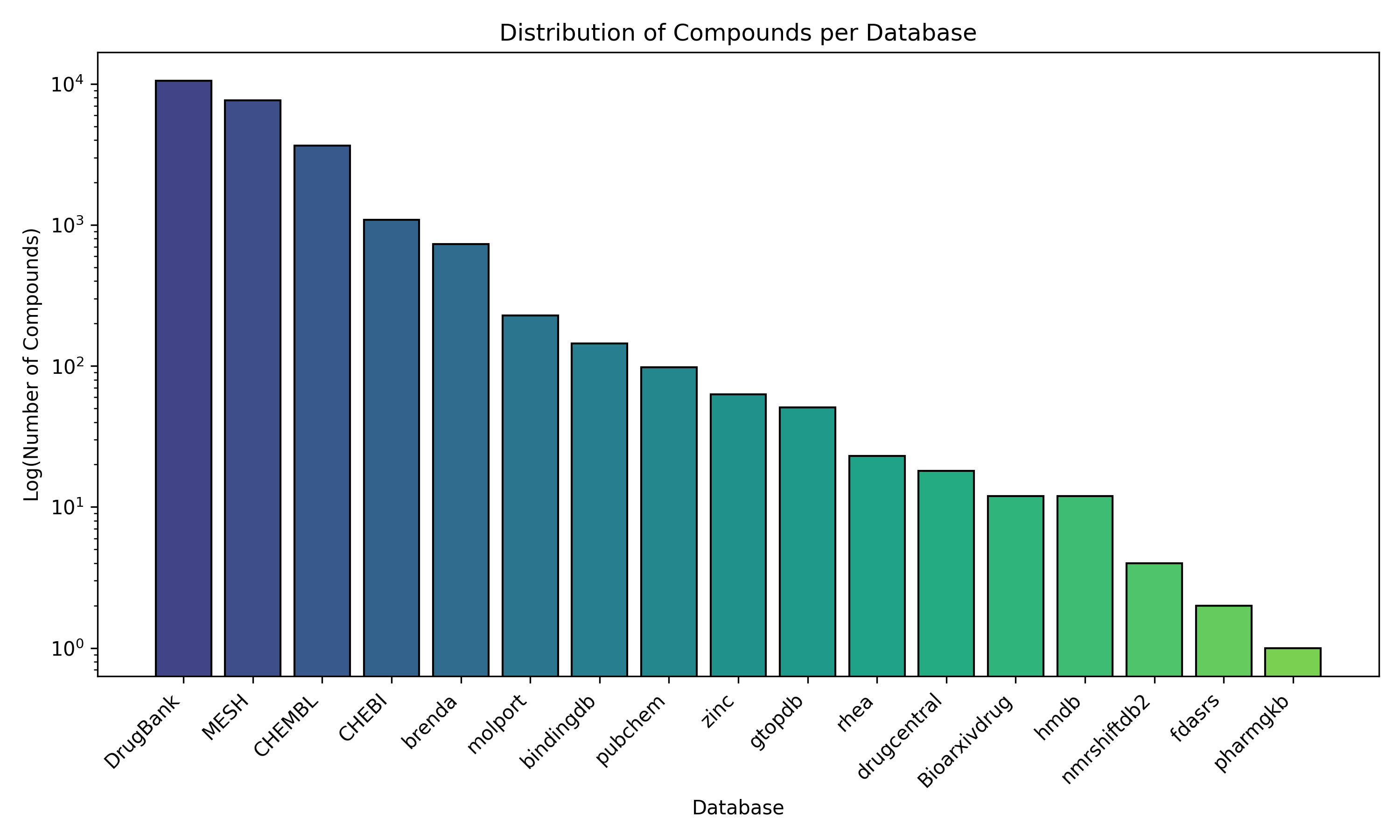
Figure 3.- Distribution of compounds per database in DRKG.
Graph neural network models
In general, GNNs represent entities and relationships in a KG as vectors in a low-dimensional space (embeddings). Then, these vectors are scored to predict new triplets. The scoring function can be based on distance or similarity measures, depending on the type of GNN. During the training process, there is a loss function that measures the difference between the predicted and the true triplets. The goal is to minimize this loss function. Also, there is a negative generator that creates false triplets to train the model. The negative generator creates triplets by replacing the subject, predicate, or object of a true triplet with a random entity of the same type.
The following figure illustrates the general structure of a knowledge graph neural network (KGNN):
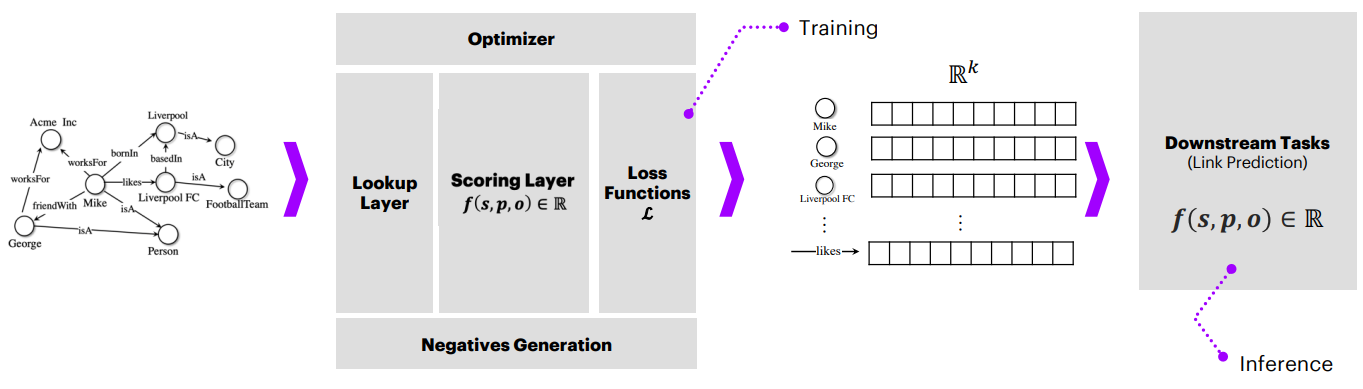
Figure 4.- Strcuture of a Knowledge Graph Neural Network. Obtained from [1].
For this project, four GNN algorithms, namely PairRE, DistMult, ERMLP, and TransR, were trained to predict new drug-disease associations using the Drug Repurposing Knowledge Graph (DRKG). These algorithms are implemented in the PyKEEN library. The models were trained using the Marging Ranking Loss function and a random seed of 1235. The rest of the hyperparameters were the default values of the library.
First, the models were trained for 50 epochs with a general evaluation procedure using all the triplets in the DRKG. In this way, the evaluation results reflected the link prediction performance for all the entity pairs in the KG. Next, new models were trained for 10 epochs with a drug repurposing evaluation procedure using only the triplets that involve drugs and diseases. Here, it was shown the link prediction performance for the task of predicting new drug-disease associations.
The trained models are available through Zenodo with the following DOI: 10.5281/zenodo.10010151.
Evaluation
The KGNN models were evaluated: a) intrinsically, within the scope of the knowledge graph and its defined triples, and b) externally, against a ground truth (drugs on clinical trials to treat dengue) to understand their predictive power over real-world information.
Internal evaluation
Two standard rank-based metrics were used to measure each KGNN model’s intrinsic performance on link prediction:
- Adjusted Mean Rank(AMR): the ratio of the Mean Rank to the Expected Mean Rank, assessing a model’s performance independently of the underlying set size. It lies on the open interval (0,2), where lower is better.
- Hits@k: the fraction of times when the correct or “true” entity appears under the top-k entities in the ranked list. The value of hits@k is between 0 and 1. The larger the value, the better the model works. For this project, I estimated hits@1, hits@3, hits@5, and hits@10 metrics
All the internal evaluation metrics were calculated using the PyKEEN library. I reported the optimistic rank values for both the tail and head entities, which assumes that the true choice is on the first position of all those with equal score when there are multiple choices. More details about how the evaluation for KGNN models works in PyKEEN can be found here.
External evaluation
To validate the KGNN models externally, I analyzed the predicted ranked compound list against the drugs on clinical trials to treat dengue defined in ground truth using the following metrics:
- First hit: the ranking position at which compounds proposed by a KGNN model match one from the ground truth database.
- Median hit: the ranking position at which compounds proposed by a KGNN model match 50% of the compounds from the ground truth database.
- Last hit: the ranking position at which compounds proposed by a KGNN model match all the compounds from the ground truth database.
For all these metrics, the smaller the value, the better, meaning that a model with lower “first”, “median”, or “last hit” values compared to another one, matches real-world compounds using fewer predictions.
The ground truth database was obtained from the ClinicalTrials.gov website. I searched for clinical trials that use drugs to treat dengue. I found 21 clinical trials that use 16 drugs to treat this disease. Also, I look for the IDs of these drugs from the 17 compound databases in the DRKG using the CHEMBL API and manual validation. The list of the drugs in the ground truth database and their IDs in the compound databases of DRKG are available in the dengue_validated_drugs_clin.csv.
Results
Internal performance evaluation results
The KGNN models were first evaluated on their general link prediction performance across the entire DRKG (trained for 50 epochs) and then specifically for the drug repurposing task (trained for 10 epochs).
The following tables show that PairRE achieved the best overall link prediction performance on the full graph. However, when focusing to drug-repurposing triplets, DistMult and ERMLP showed higher values, suggesting that while some models excel at capturing global graph topology, others are better suited for specific biomedical associations.
Table 1. Internal evaluation for all triplets (50 epochs)
| Model | Adjusted Mean Rank (AMR) | Hits@1 | Hits@3 | Hits@5 | Hits@10 |
|---|---|---|---|---|---|
| PairRE | 0.0179 | 0.0211 | 0.114 | 0.160 | 0.221 |
| ERMLP | 0.0188 | 0.0276 | 0.079 | 0.110 | 0.164 |
| TransR | 0.0193 | 0.0121 | 0.064 | 0.088 | 0.132 |
| DistMult | 0.0400 | 0.0160 | 0.036 | 0.050 | 0.076 |
Table 2. Internal evaluation for drug repurposing triplets (10 epochs)
| Model | Adjusted Mean Rank (AMR) | Hits@1 | Hits@3 | Hits@5 | Hits@10 |
|---|---|---|---|---|---|
| DistMult | 0.0293 | 0.0078 | 0.0188 | 0.0270 | 0.0424 |
| ERMLP | 0.0310 | 0.0086 | 0.0209 | 0.0286 | 0.0461 |
| PairRE | 0.0333 | 0.0041 | 0.0119 | 0.0178 | 0.0314 |
| TransR | 0.0392 | 0.0013 | 0.0042 | 0.0073 | 0.0139 |
External performance evaluation results
The following tables show how well the models ranked the 16 drugs known to be in clinical trials for Dengue (ground truth).
Table 3. External evaluation for all triplets
| Model | First Hit | Median Hit | Last Hit |
|---|---|---|---|
| ERMLP | 26 | 1,432 | 11,354 |
| DistMult | 31 | 1,407 | 10,689 |
| PairRE | 219 | 1,397 | 17,149 |
| TransR | 440 | 2,297 | 16,402 |
Table 4. External evaluation for drug repurposing triplets
| Model | First Hit | Median Hit | Last Hit |
|---|---|---|---|
| ERMLP | 9 | 2,849 | 15,369 |
| DistMult | 17 | 2,655 | 18,005 |
| PairRE | 82 | 1,352 | 5,619 |
| TransR | 323 | 4,321 | 18,393 |
The ERMLP model demonstrated the strongest predictive power for real-world applications, achieving a “First Hit” at the 9th position in the drug repurposing task. This indicates that the model successfully ranked a clinically validated drug within its top 10 predictions, outperforming more complex models like TransR in identifying relevant therapeutic candidates.
Predicted repurposed drugs for Dengue
Based on the ERMLP model, the top three predictions for potential repurposed drugs are:
- Betamethasone: A corticosteroid with anti-inflammatory and immunosuppressive properties.
- Dexamethasone: A potent glucocorticoid often used to treat severe inflammation.
- Aspirin: A common nonsteroidal anti-inflammatory drug (NSAID).
The identification of these compounds suggests that the models effectively captured the importance of managing the systemic inflammatory response and pain symptoms characteristic of Dengue infection, as all top candidates are established anti-inflammatory or analgesic agents. However, this focus on symptomatic relief suggests the models may be biased toward well-documented inflammatory pathways in the KG, potentially overlooking compounds with direct antiviral mechanisms against the dengue virus.
Additional information
More details about the biological background of the project, the exploratory data analysis, feature engineering, models training, hyperparameter tuning, interpretation of the results, and ideas for further work are available in the GitHub repository of this project and its presentation.
Citation
Ayala-Ruano, S. (2023). DengueDrugRep: Drug repurposing for dengue using a biomedicine knowledge graph and graph neural networks (Version 1.0.0) [Model]. Zenodo. doi: doi.org/10.5281/zenodo.10010151.