Source code Manuscript
I developed this project for my capstone project of the Machine learning course from the MSc in Systems Biology at Maastricht University. It was an end-to-end assignment with data pre-processing, feature engineering, training, hyperparameter tuning, and interpretation of the results.
Background
Vision is one of the main sensory pathways that enable living organisms to perceive external stimuli and interpret the world. The human visual system (HVS) involves a complex interplay between the eyes, the brain, and multiple neural pathways. To study the HVS, investigators employ various experimental methods such as psychophysics, eye tracking, and neuroimaging techniques like functional magnetic resonance imaging (fMRI). fMRI uses magnetic resonance imaging to measure the blood-oxygen-level-dependent (BOLD) variation induced by neuronal activity.
The data acquired through fMRI experiments are stored in the form of voxels, which are three-dimensional units representing tiny volume elements. Each voxel encompasses millions of brain cells, collectively forming areas with different functional properties, known as regions of interest (ROIs). Computational models are essential tools for interpreting the large datasets produced by fMRI experiments.
The goal of this project is to employ machine learning techniques to forecast the neural visual responses triggered by naturalistic scenes. These computational models aim to replicate the process through which neuronal activity encodes to visual stimuli aroused by the external environment. The following figure gives an schematic representation of the brain encoding and decoding processes.
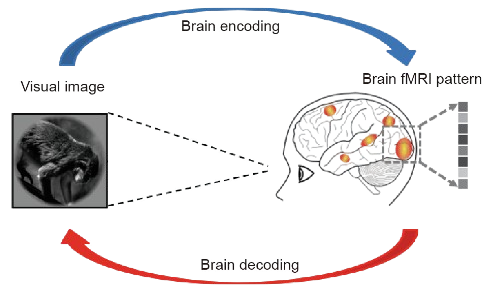
Figure 1.- Brain encoding and decoding in fMRI. Obtained from [1].
Visual encoding models based on fMRI data employ algorithms that transform image pixels into model features and map these features to brain activity. This framework enables the prediction of neural responses from images. The following figure illustrates the mapping between the pixel, feature, and brain spaces.
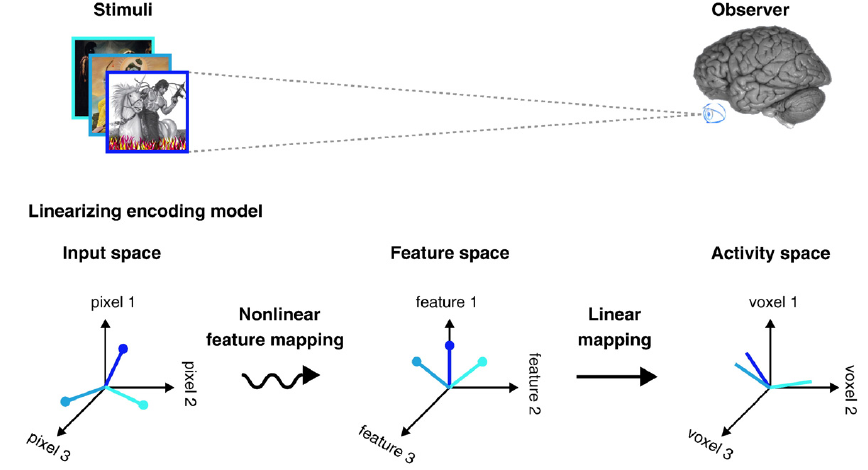
Figure 2.- The general architecture of visual encoding models that consists of three spaces (the input space, the feature space, and the brain activity space) and two in-between mappings. Obtained from [2].
Dataset
The data for this project is part of the [Natural Scenes Dataset][nsd] (NSD), a massive dataset of 7T fMRI responses to images of natural scenes coming from the [COCO dataset][coco]. The training dataset consists of brain responses measured at 10.000 brain locations (voxels) to 8857 images (in jpg format) for one subject. The 10.000 voxels are distributed around the visual pathway and may encode perceptual and semantic features in different proportions. The test dataset comprises 984 images (in jpg format), and the goal is to predict the brain responses to these images.
You can access the dataset through Zenodo with the following DOI: doi.org/10.5281/zenodo.7979729.
The training dataset was split into training and validation partitions with an 80/20 ratio. The training partition was used to train the models, and the validation partition was used to evaluate the models. The test dataset was used to make predictions with the best model on unseen data.
The figure below summarizes the data splitting process and the rest of the steps to build the visual encoding models.
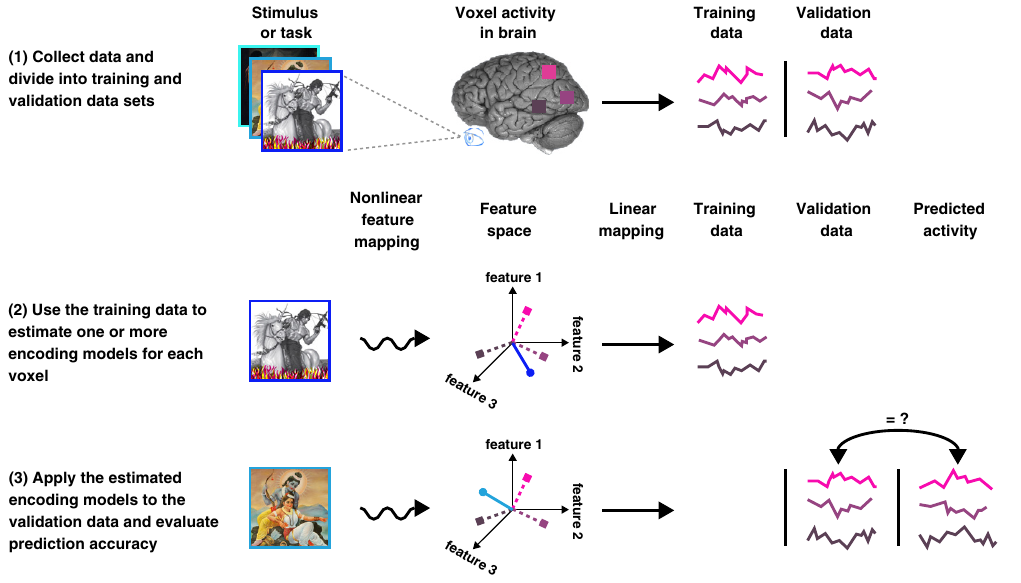
Figure 3.- Summary of the main steps for the creation of the visual encoding models. Obtained from [2].
Feature engineering
Due to the high dimensionality of the feature representation of images using the raw pixel values (i.e., the original images have a size of 425x425 and 3 channels (RGB), which results in a feature representation of 425x425x3 = 541875 features), I used the representations obtained from different layers of pretrained CNNs to obtain a lower dimensional representation of the images. In this case, I tried various layers of four different pretrained CNNs: AlexNet, VGG16, ResNet50, and InceptionV3, available in the torchvision package.
A graphical representation of the feature engineering process is presented below.
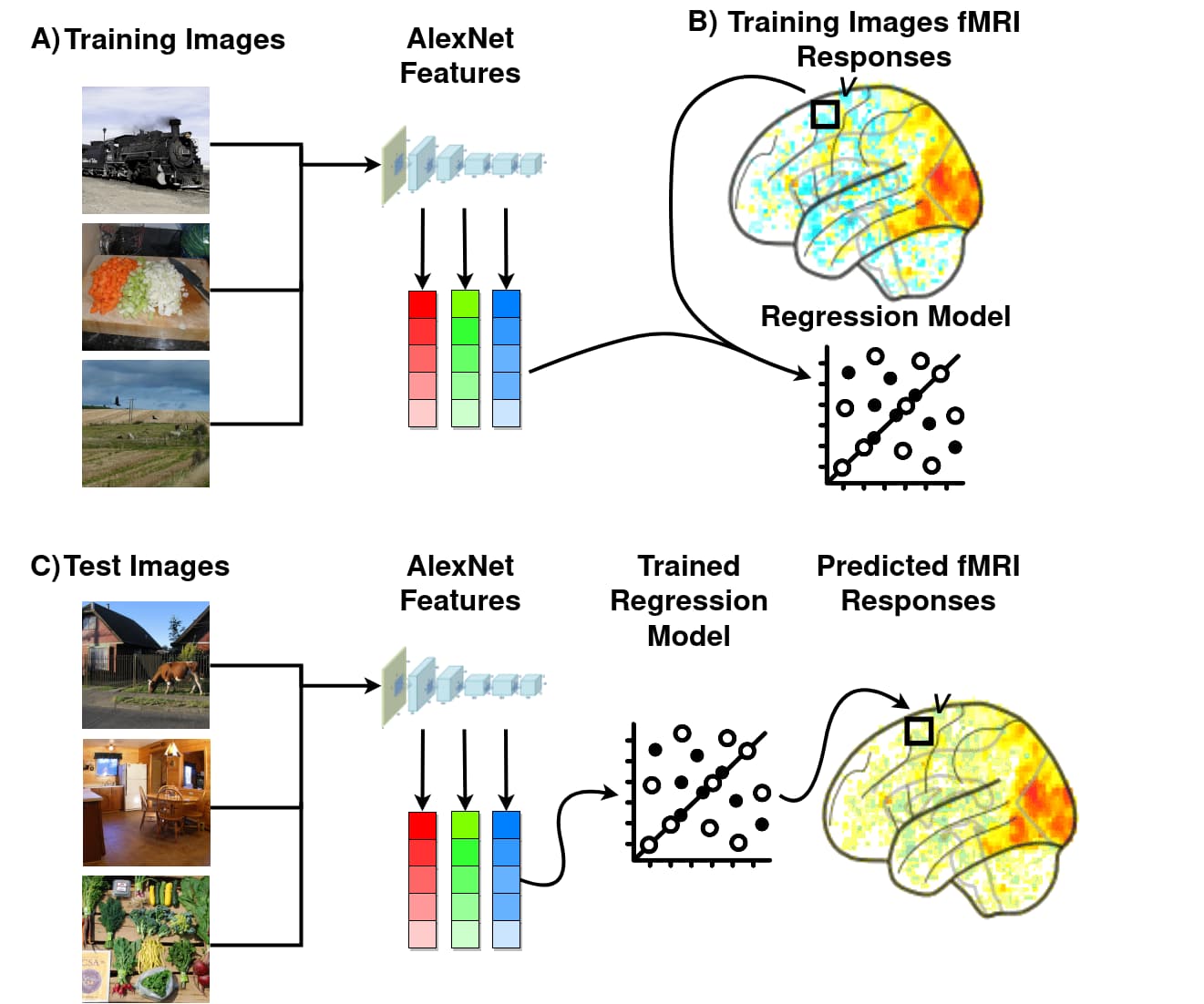
Figure 4.- Diagram of the feature engineering stage. Obtained from [3].
The feature representations of the images was obtained by passing the images through the pretrained CNNs and extracting the output of the desired layer. The size of the feature vectors at this point was still very large, so I used PCA to overcome this problem and got a set of 30 features. I fit the PCA on the training images features, and used it to downsample the training, validation and test images features.
I evaluated the best feature representation by training a simple linear regression model to predict the brain activity of the voxels from the feature representation of the images. The best feature representation was the one that resulted in the highest encoding accuracy (i.e., median correlation between the predicted and actual brain activity of the voxels) on the validation set.
Machine learning models
I trained 6 different machine learnning algorithms (linear regression - base model, ridge regression, lasso regression, elasticnet regression, k-nearest neighbours regressor, and decision tree regressor) to predict the brain activity of the voxels from the feature representation of the images. In this project, the learning task was a multioutput regression problem, where the input is the feature representation of the images and the output is the brain activity of all the voxels. Each regressor maps from the feature space to each voxel, so there is a separate encoding model per voxel, leading to voxelwise encoding models. Therefore, every model trained with this dataset have 10.000 independently regression models with n coeficients each (the number of features). As in the previous section, the best model was the one that resulted in the highest encoding accuracy on the validation set.
Results of the ML models
The best model was Lasso regression with an encoding accuracy of 0.2417 on the validation set. The best hyperparameters were alpha=0.01 and the default max_iter=1000. This model was trained with the feature representation of the images obtained from layer features.12 of the AlexNet CNN, reduced to 100 features using PCA.
| Machine Learning Model | Encoding Accuracy |
|---|---|
| Lasso (alpha=0.01) | 0.2417 |
| ElasticNet (alpha=0.001) | 0.2415 |
| Ridge (alpha=1.0) | 0.2412 |
| Linear Regression | 0.2402 |
| K-Nearest Neighbors | 0.1021 |
| Decision Tree | 0.0382 |
While the overall encoding accuracy of the best model is low, the distribution of predictions across voxels reveals important patterns: regularized linear models show right-skewed accuracy distributions with a heavy tail between 0.4-0.7, indicating that the models make accurate predictions on a subset of voxels while struggling with others. The lack of ROI information prevents us from identifying which visual areas correspond to high or low predictions.
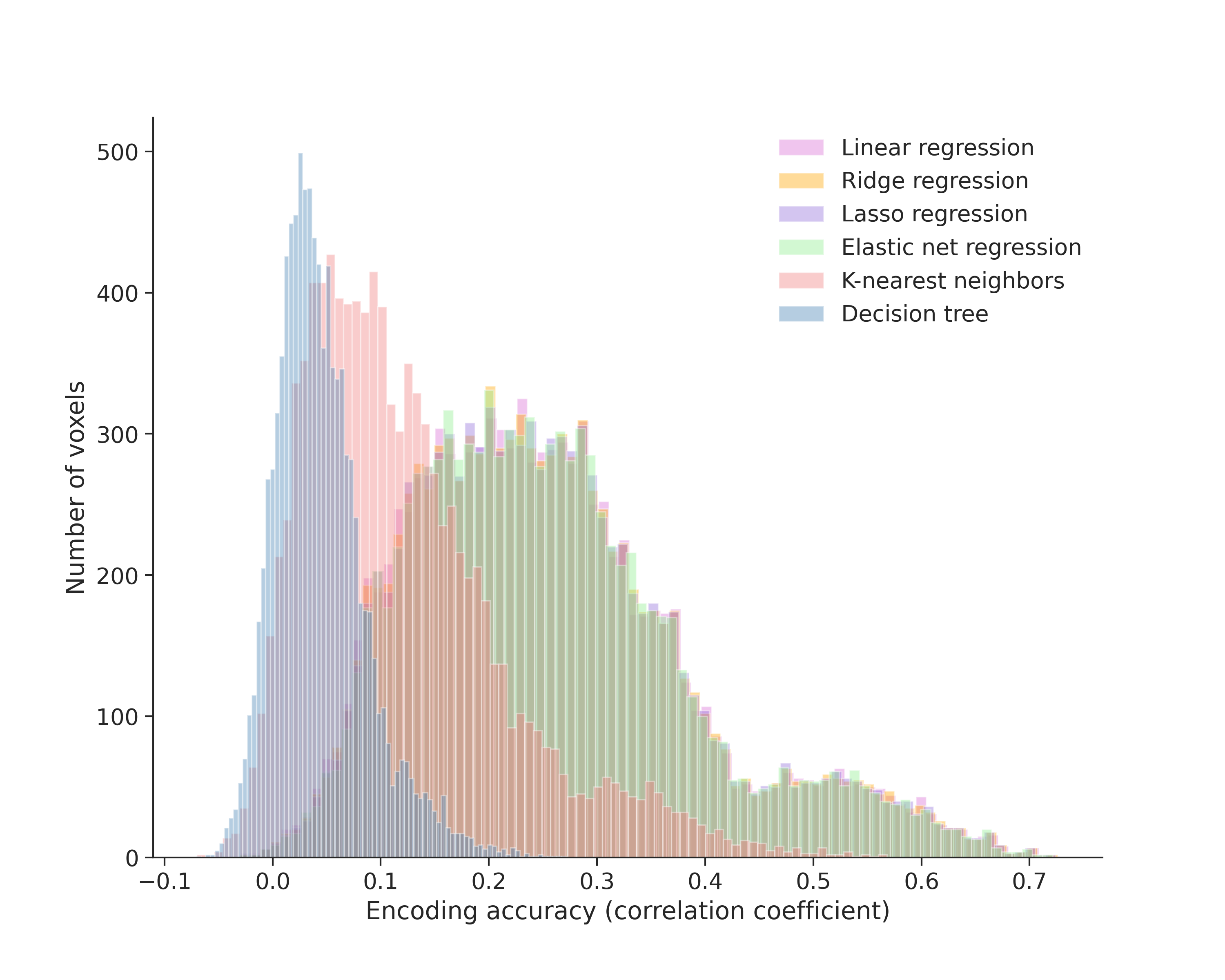
Figure 5.- Histograms of encoding accuracy for machine learning models across all voxels. Models were trained to predict neural responses to visual stimuli from naturalistic images.
To validate these findings, we compared predictions from the best and worst performing voxels. The top-performing voxel showed strong agreement between predicted and actual BOLD signals with high positive correlation, while the lowest-performing voxel showed no meaningful overlap or correlation pattern. This demonstrates that the model successfully captures neural responses for a subset of voxels but fails to generalize across the entire brain.
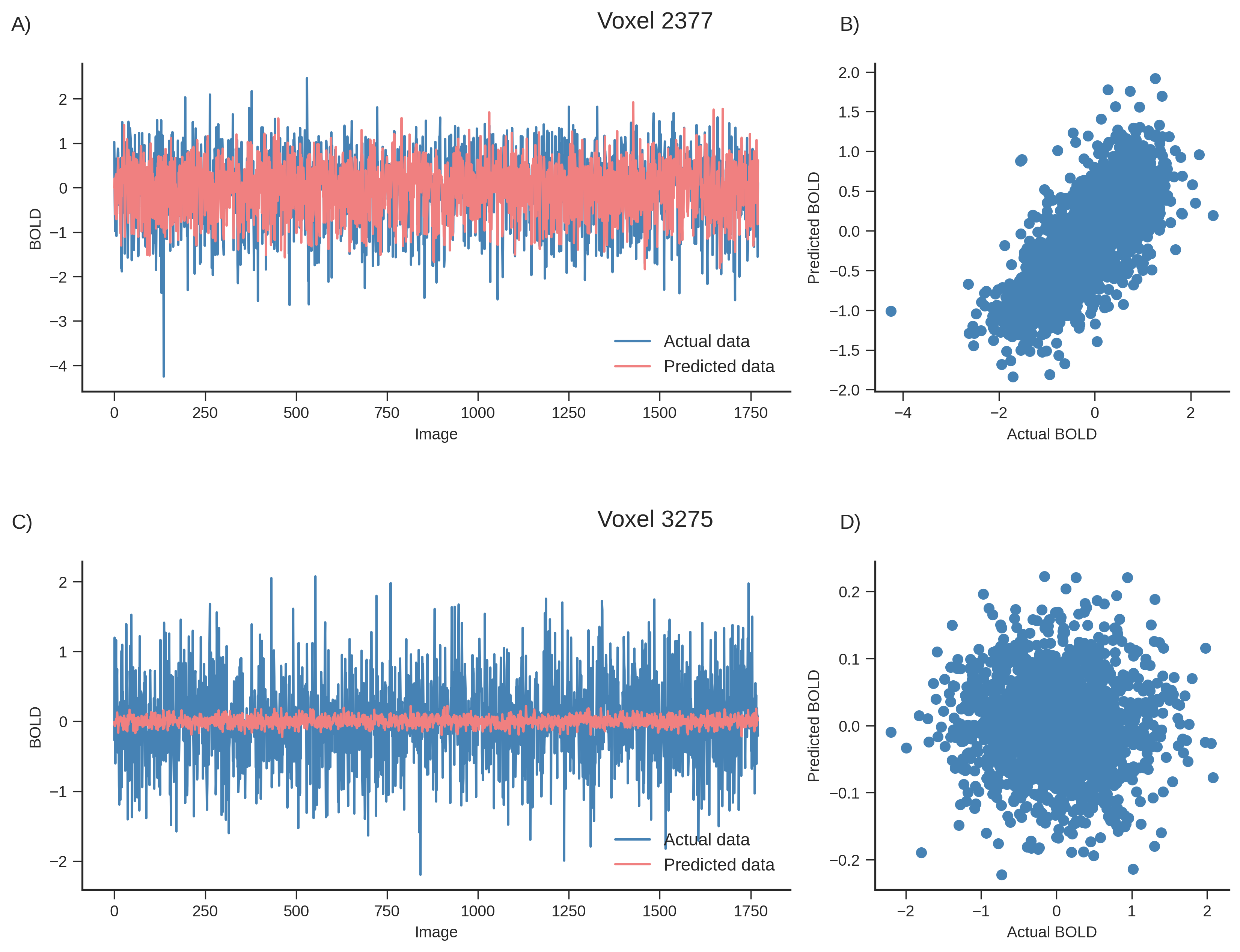
Figure 6.- Predicted vs. actual BOLD variation for the best (A-B) and worst (C-D) performing voxels from the Lasso regression model.
Additional information
More details about the biological background of the project, the exploratory data analysis, feature engineering, model training, hyperparameter tuning, interpretation of the results, and ideas for further work are available in the GitHub repository of this project and in the manuscript.
Citation
Ayala-Ruano, S. (2023). Img2brain: Predicting the neural responses to visual stimuli of naturalistic scenes using machine learning (Version 1.0.0) [Dataset/Software]. Zenodo. doi: doi.org/10.5281/zenodo.7979729.