I developed this project for my capstone project of the Network Biology course from the MSc in Systems Biology at Maastricht University. It was an end-to-end assignment with data pre-processing, network inference and analysis, and interpretation of the results.

Background
The Earth’s microbiome - all microbial communities, their structural components, and metabolic byproducts - constitutes the largest portion of biodiversity on our planet. Due to their widespread presence, microbes profoundly impact the biosphere, collectively regulating the global biogeochemical cycles, soil fertility, and other processes. In particular, it is crucial to study the soil microbial diversity of tropical ecosystems facing threats due to extensive deforestation for crop cultivation or pasture expansion. While standard metagenomic analyses identify taxonomic composition, they often overlook ecological interactions within microbial communities.
Microbial association networks (MANs) provide insights into potential interactions among microbes, including mutualism, competition, and more. Furthermore, these networks can reveal communities that share ecological functions or keystone taxa playing crucial roles in the system. In MANs, nodes correspond to Operational Taxonomical Units (OTUs) at a given taxonomic rank, and edges between nodes denote significant co-presence (positive relationships) or mutual exclusion (negative relationships) patterns in OTU abundances across samples.
The figure below illustrates the workflow to infer MANs from meta-omics datasets.
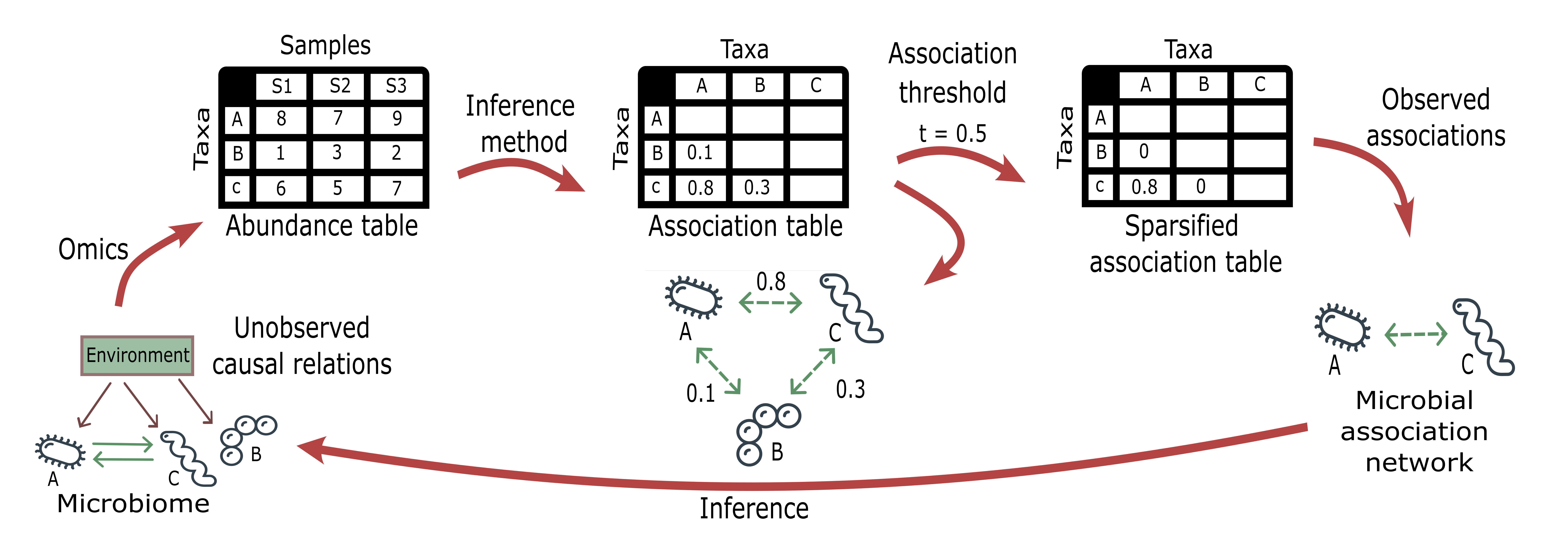
Figure 1.- Workflow to infer microbial association networks (MANs) from meta-omics datasets.
In this project, the CCLasso (correlation-based) and SPRING (conditional dependence-based) methods were used to explore differences in microbiomes found in the rainforest and converted pasturelands of the northwest Colombian Amazon region. The NetCoMi R package v1.1 was the main framework to perform the network inference, analysis, visualization, and comparison.
Dataset
The raw sequencing data used for this project is available at the PRJEB44163 project from the European Nucleotide Archive (ENA) database. This research collected 52 soil samples from the Colombian Amazon region: 36 from rainforest areas and 16 from converted pasturelands. The rainforest samples were the reference with minimal intervention, while the pastureland represented the land use systems.
MGnify is a platform that automates the analysis of metagenomics datasets from ENA and other databases. The abundance table and taxonomic profiles to infer the MANs in this project were obtained from the MGYS00005779 study.
The first step was downloading the data and metadata using the MGnifyR package v0.1, enabling the use of the MGnify API in R scripts. Then, the data was preprocessed and manipulated using the Phyloseq v1.44 and Microbiome v1.22 R packages. The dataset was filtered to retrieve only bacterial OTUs, excluding taxa from other life kingdoms. In addition, the data was aggregated at the family taxonomic level, obtaining 200 OTUs.
Results
1. Comparison of inference methods
The initial stage of the project involved a comparison between CCLasso and SPRING. CCLasso was found to have more favorable topological features for capturing microbiome interactions, including more efficient information flow (lower average path length) and clearer identification of degree-based hub nodes.
The following table summarizes the topological features of all networks:
| Metric | Forest SPRING | Forest CCLasso | Pasture SPRING | Pasture CCLasso |
|---|---|---|---|---|
| Nodes | 108 | 43 | 95 | 56 |
| Edges | 183 | 108 | 185 | 113 |
| Modularity | 0.69 | 0.46 | 0.71 | 0.43 |
| Avg. Path Length | 6.02 | 3.20 | 6.96 | 2.78 |
2. Impact of land use conversion
The visual analysis and statistical comparison via non-parametric permutation testing confirmed a significant increase in negative interactions in pastureland networks compared to forest counterparts. This suggests a shift toward antagonistic relationships, such as increased competition and predation, following land perturbation. The figures below show the forest and pasturland networks inferred with CCLAsso and SPRING.
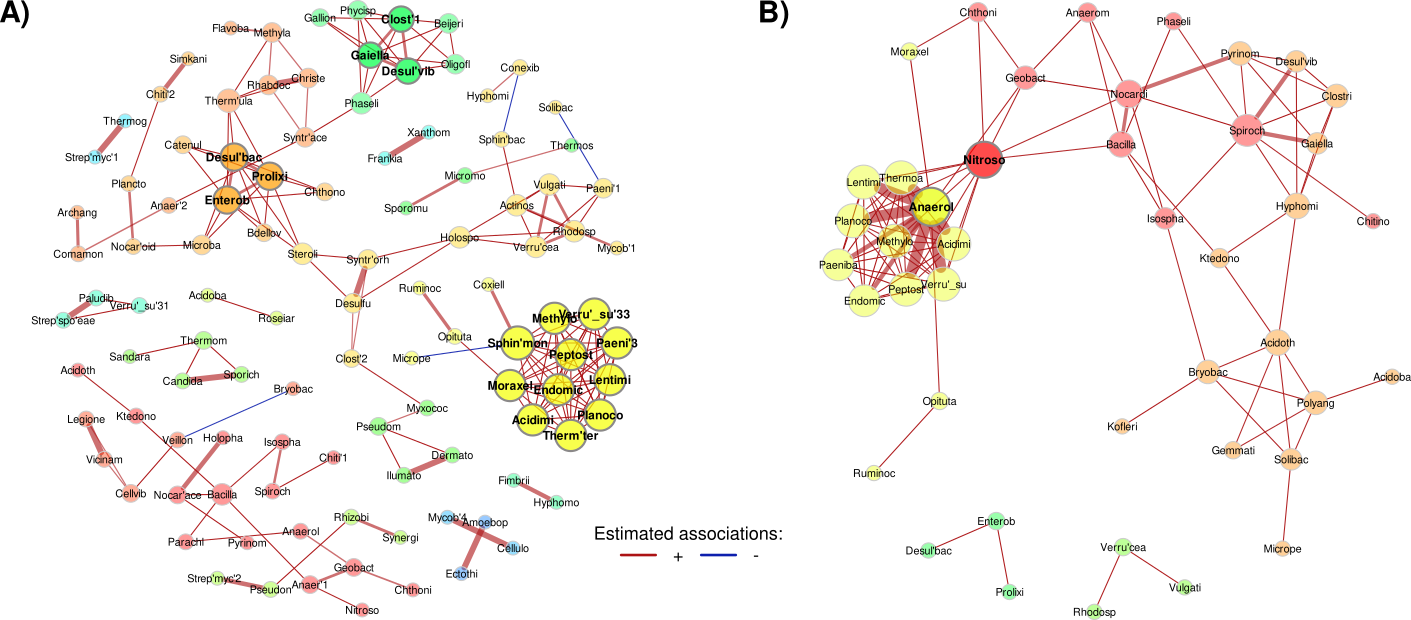
Figure 2.- Forest bacterial association networks inferred with A) SPRING and B) CCLasso. Nodes represent family-level OTUs, colored by their community and sized by degree.
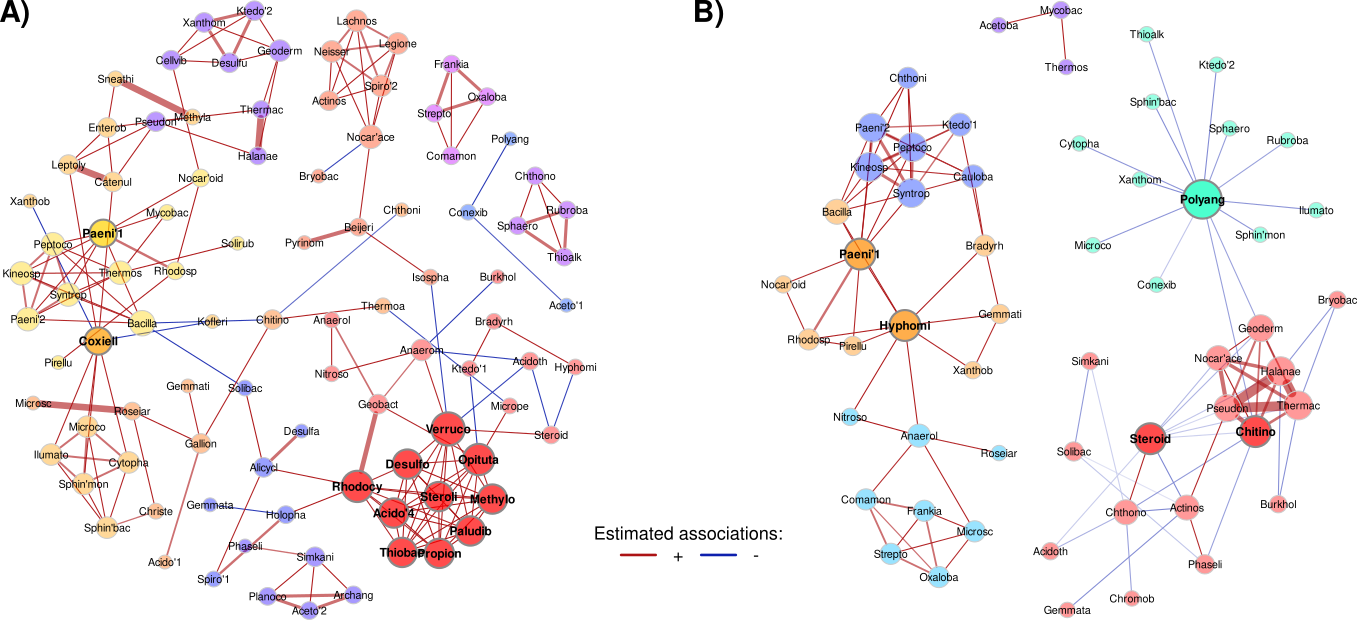
Figure 3.- Pasture bacterial association networks inferred with A) SPRING and B) CCLasso. Nodes represent family-level OTUs, colored by their community and sized by degree. Note the increase in negative associations (blue edges).
3. Taxonomic shifts by Phylum
The land conversion resulted in these taxonomic patterns:
- Acidobacteria: Significant decrease in pasturelands, likely due to increased soil pH and carbon.
- Actinobacteria: Increase in pasturelands, potentially linked to their role in competitive ecological interactions and antibiotic production.
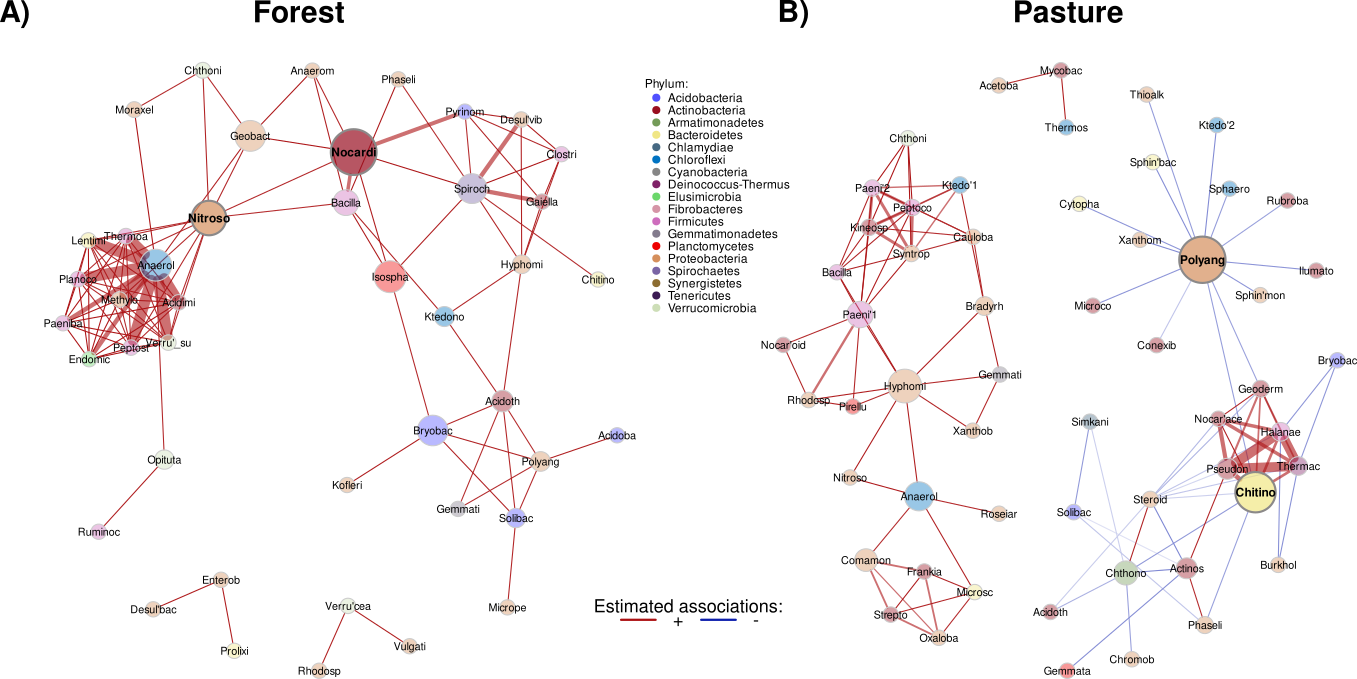
Figure 4.- Figure 4. A) Forest and B) Pasture networks inferred with CCLasso. Nodes represent family-level OTUs, colored by their phylum and sized by betweenness. Red edges correspond to positive associations and blue edges to negative ones.
Additional information
More information about the microbiome analysis, a detailed description of the methodology, an in-depth discussion of the results, and further details are available on the GitHub repository of this project and in the manuscript.