Publication Source code
I contributed to this project as part of my work as research assistant at the Applied Signal Processing and Machine Learning Group and the Grupo de Medicina Molecular y Traslacional - Universidad San Francisco de Quito, in Ecuador.
Summary
Antimicrobial peptides (AMPs) are small bioactive chemicals that have appeared as promising compounds to treat a wide range of diseases. The effectiveness of AMPs resides in the wide range of mechanisms they can use for both killing microbes and modulating immune responses. However, the AMPs’ chemical space (AMPCS) is huge, it is estimated that there exist more than 10^65 unique sequences of peptides with 50 residues or fewer, which represent a big challenge for the discovery of new promising sequences and the identification of common features, motifs, or relevant biological functions shared by these peptides.
Therefore, we developed StarPep toolbox, an open-source software to study the AMPCS using networks, clustering, and similarity-searching models, which can contribute to peptide drug repurposing, development, and optimization.

This tool was developed as a Java desktop application that integrates the functionalities of several open-source projects. The graphical user interface was built on top of the NetBeans Platform, using the Java SE Runtime Environment 8. The graph database structure was implemented with the Neo4j platform. Some visualization features and the calculation of network properties were based on Gephi. The sequence alignment algorithms were implemented using the BioJava API.
The AMPs were collected from a large variety of biological data sources to be organized into an integrated graph database called starPepDB, composed of 45.120 AMPs and their metadata. This integrated graph database is embedded into StarPep toolbox to enable end-user querying, filtering, visualizing, and analyzing the AMPs, taking advantage of network-based representations.
The main modules of StarPep toolbox are listed below:
AMPs’ chemical space filtering: Obtain a subset of AMPs from the StarPepDB using their metadata (function, target pathogen, biological origin, chemical modifications, original database, and cross-referenced entries to PDB, PubMed, and UniProt).
Molecular descriptors: Calculate molecular descriptors of the AMPs by applying statistical and aggregation operators on physicochemical amino acid properties (e.g., net charge, isoelectric point, molecular weight, etc.).
Network Science: Build different types of networks (metadata, chemical space, and half-space proximal) and calculate global/local properties, centrality metrics, communities, among other metrics.
Similarity searching: Create multi-query similarity searching models that can lead to the repurposing of AMPs with novel functional activities.
The figure below shows the software architecture and components of StarPep toolbox:
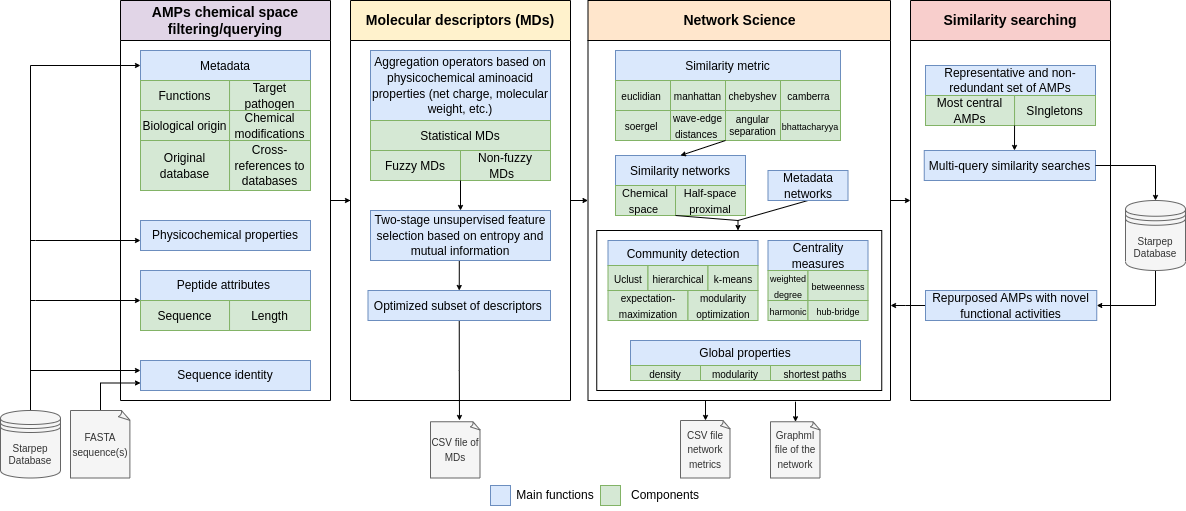
Figure 1.- StarPep modules and components. The software has four main modules to load and filter AMPs, calculate MDs, create and analyze networks, and develop multi-query similarity-searching models.
For more details about the software, read its online User Guide and publication. Also, you can check the source code to explore the implementation of the software.
Citation
Aguilera-Mendoza, L., Ayala-Ruano, S.^, Martinez-Rios, F., Chavez, E., García-Jacas, C. R., Brizuela, C. A., & Marrero-Ponce, Y. (2023). StarPep Toolbox: an open-source software to assist chemical space analysis of bioactive peptides and their functions using complex networks. Bioinformatics, 39 (8), btad506. doi: doi.org/10.1093/bioinformatics/btad506.
^co-first author