Publication Source code
I contributed to this project as part of my work as research assistant at the Multiomics Network Analytics Group (MoNA) - Novo Nordisk Foundation Center for Biosustainability (DTU Biosustain).

Summary
VueGen is a tool that automates the creation of reports from bioinformatics outputs, allowing researchers with minimal coding experience to communicate their results effectively. With VueGen, users can produce reports by simply specifying a directory containing output files, such as plots, tables, networks, Markdown text, HTML components, and API calls, along with the report format. Supported formats include documents (PDF, HTML, DOCX, ODT), presentations (PPTX, Reveal.js), Jupyter notebooks, and Streamlit web applications. By generating reports in all these formats, VueGen simplifies the reporting process, eliminating the need to manage multiple pipelines or manually reformat content.
A YAML configuration file is generated from the directory to define the structure of the report. Users can customize the report by modifying the configuration file, or they can create their own configuration file instead of passing a directory as input. The configuration file specifies the structure of the report, including sections, subsections, and various components such as plots, dataframes, markdown, html, and API calls.
An overview of the VueGen workflow is shown in the figure below:
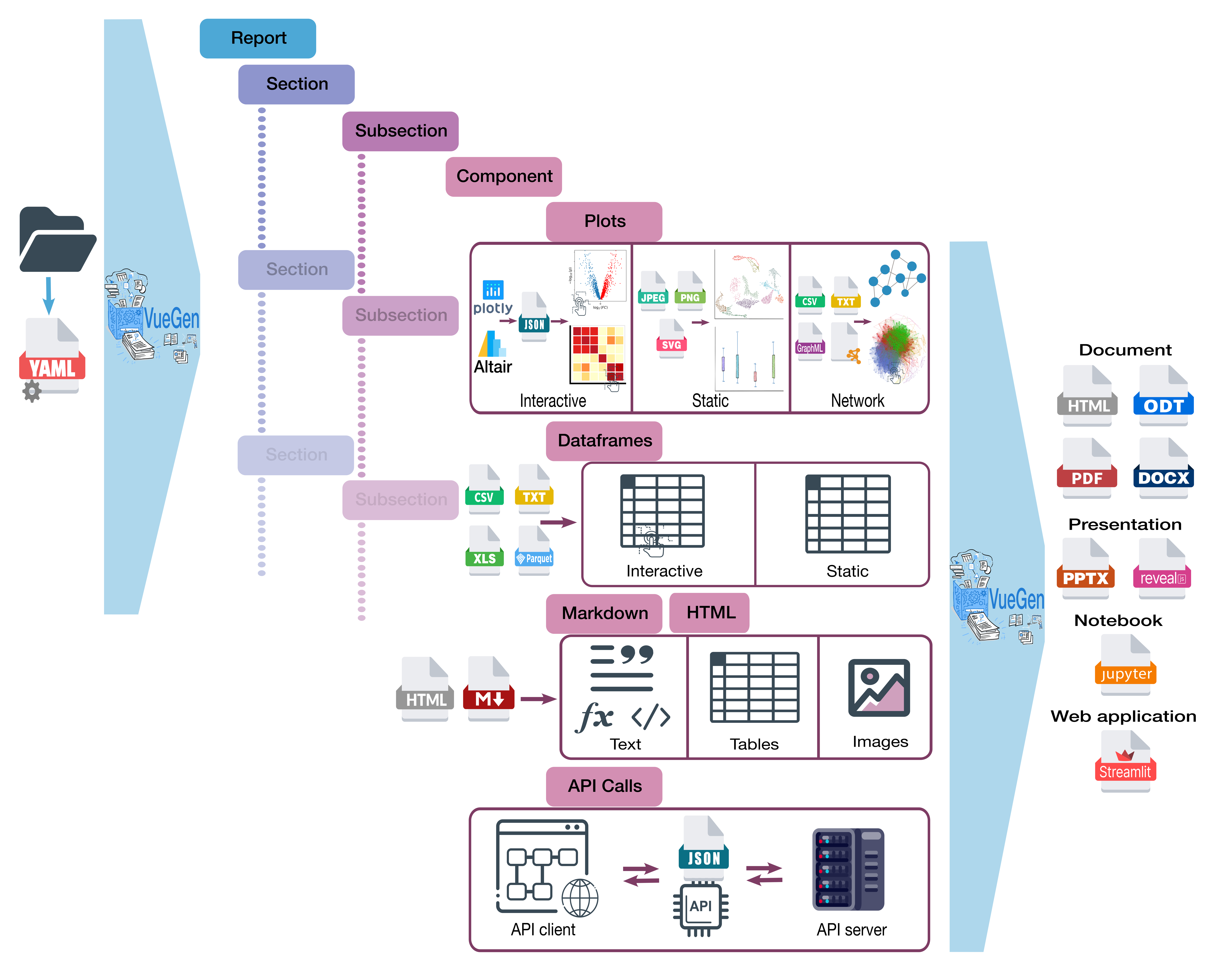
Figure 1.- Graphical overview of VueGen’s workflow and components. The diagram illustrates VueGen’s pipeline and report structure, starting from either a directory or a YAML configuration file.
Its modular and cross-platform design facilitates integration into various analytical workflows and enables researchers with minimal or no coding experience to communicate their results effectively. By combining ease of use for non-programmers with extensibility for advanced users, VueGen benefits a broad audience and shifts the focus from time-consuming report generation to the meaningful analysis and interpretation of results.
A schema diagram was created to illustrate the structure of the configuration file and the relationships between its elements:
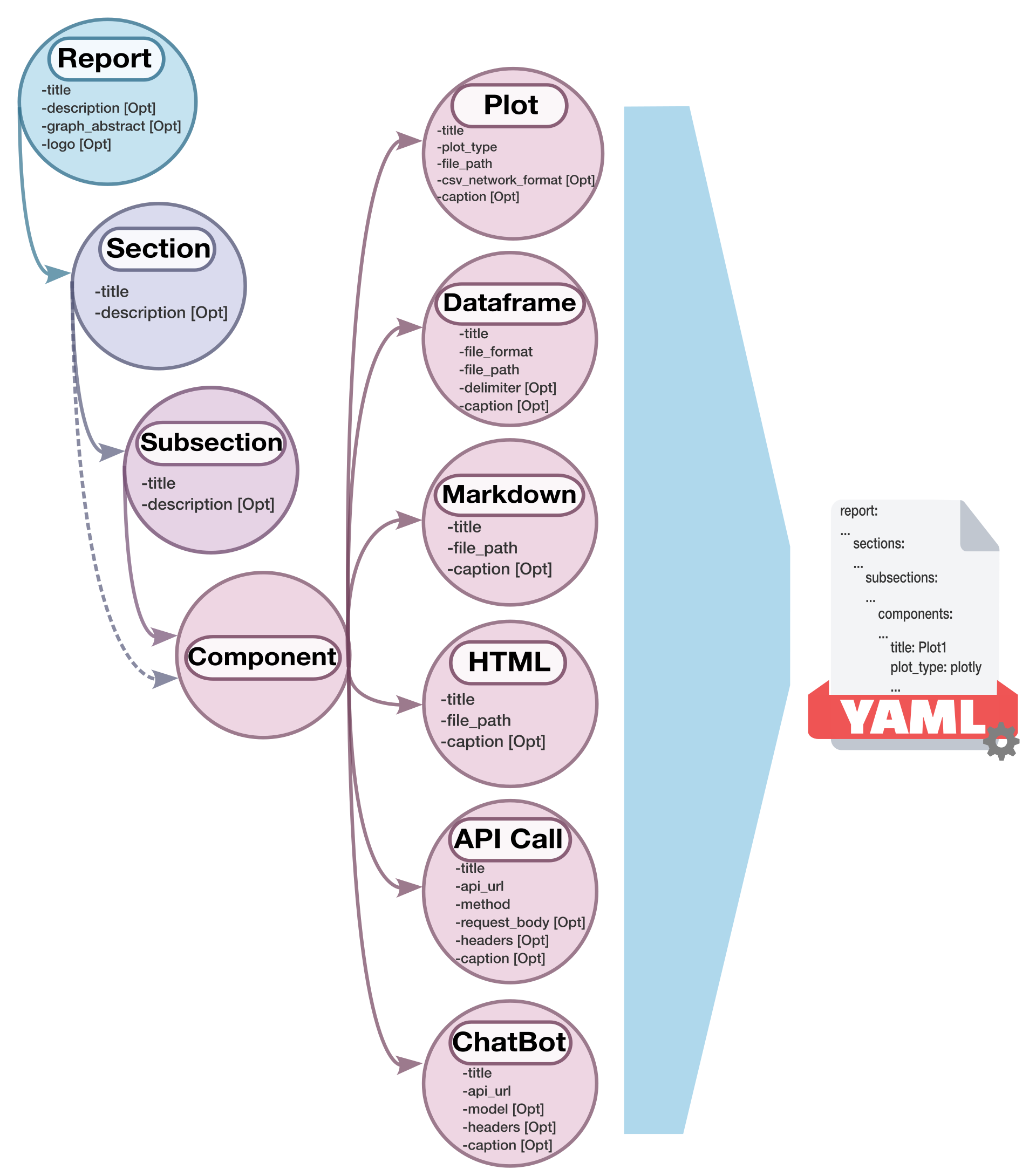
Figure 2.- VueGen schema diagram (v0.4.0). Circles represent elements of the report configuration, including report, sections, subsections, and components. Arrows indicate hierarchical relationships and data flow within the configuration file.
Also, the class diagram for the project’s current version is presented below to show the architecture and relationships between classes:
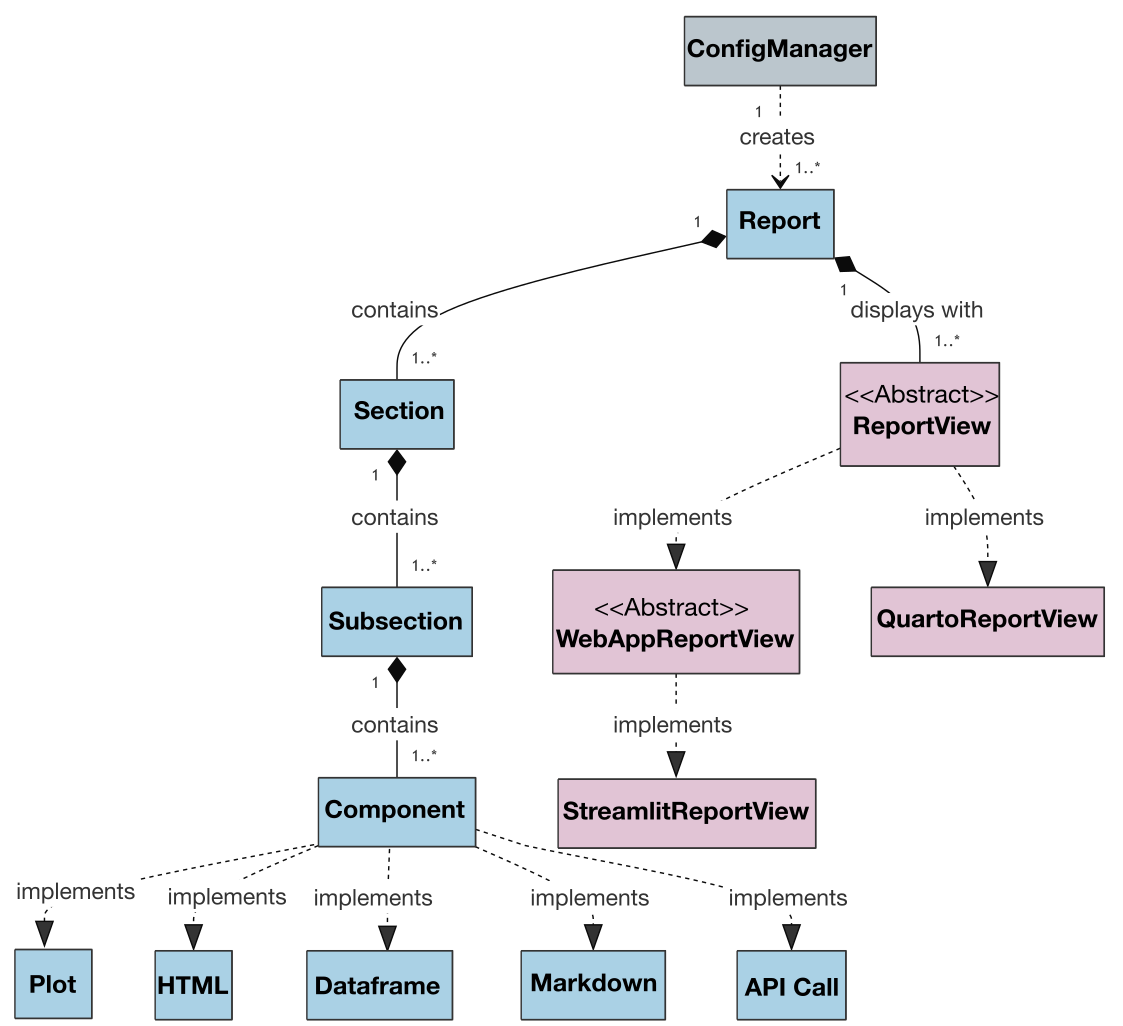
Figure 3.- VueGen class diagram (v0.4.0). Boxes represent classes, and arrows show their relationships. Colors indicate functionality: gray for classes managing input files and metadata, blue for those defining the report structure, and red for classes generating reports.
Implementation
VueGen offers various implementation options for both non-technical and experienced users. It is available as a Python package, Docker image, nf-core module, and cross-platform desktop application with a user-friendly interface, making it accessible and customizable for different user needs and expertise levels. To streamline software maintenance, VueGen applies a continuous integration and deployment (CI/CD) pipeline for automated releases of the tool and its documentation.
Additional information
The VueGen documentation is available at vuegen.readthedocs.io, where you can find detailed information of the package’s classes and functions, installation and execution instructions, and case studies to demonstrate its functionality. The publication describing the software is available in Bioinformatics Advances. The source code is openly available on GitHub, where you can also find the latest updates and contribute to the project.
Citation
Ayala-Ruano, S., Webel, H., & Santos, A. (2025). VueGen: Automating the generation of scientific reports. Bioinformatics Advances, vbaf149. doi: doi.org/10.1093/bioadv/vbaf149.